Winkelmass means “angle measure” in German. In forestry the term has been adopted to refer to a specific metric used to describe the regularity of tree spatial distribution. Literature on the winkelmass traces mostly to Klaus von Gadow, a researcher split between the University of Göttingen and Beijing Forestry University. The winkelmass is quite simple at it’s core. For each tree, a given number of nearest neighbours are identified, usually four. Then, going clockwise around the focal tree, the angle between each neighbour and the next neighbour is calculated. For each angle, if the angle is less than the ‘critical angle’ i.e. 360 degrees divided by the number of neighbours considered, a value of 1 is recorded, otherwise, a value of 0 is recorded. Finally, the sum of these values divided by the number of neighbours gives the winkelmass. In LaTeX math:
W_{i} = \frac{1}{k} \sum_{j=1}^{k} v_{j}
where W is the winkelmass, k is the number of neighbours considered, and v_{j} is either 0 or 1 depending on the angle between two neighbours.
To calculate the plot-level winkelmass, simply take the mean of all W_{i} values in the plot.
I wrote a function in R to calculate the Winkelmass for all trees in a plot:
#' Winkelmass (spatial regularity of trees)
#'
#' @param x vector of individual x axis coordinates
#' @param y vector of individual y axis coordinates
#' @param k number of neighbours to consider
#'
#' @references von Gadow, K., Hui, G. Y. (2001). Characterising forest spatial
#' structure and diversity. Sustainable Forestry in Temperate Regions. Proc. of
#' an international workshop organized at the University of Lund, Sweden.
#' Pages 20- 30.
#'
winkelmass <- function(x, y, k = 4) {
dat_sf <- sf::st_as_sf(data.frame(x,y), coords = c("x", "y"))
dists <- suppressMessages(nngeo::st_nn(dat_sf, dat_sf, k = k+1,
progress = FALSE))
a0 <- 360 / k
wi <- unlist(lapply(dists, function(i) {
focal_sfg <- sf::st_geometry(dat_sf[i[1],])[[1]]
nb_sfg <- sf::st_geometry(dat_sf[i[-1],])
nb_angles <- sort(unlist(lapply(nb_sfg, function(j) {
angleCalc(focal_sfg, j)
})))
aj <- nb_angles - c(NA, head(nb_angles, -1))
aj[1] <- nb_angles[k] - nb_angles[1]
aj <- ifelse(aj > 180, 360 - aj, aj)
aj <- round(aj, 1)
sum(aj < a0)
}))
out <- 1 / k * wi
return(out)
}
The function uses {nngeo} to find nearest neighbours of each tree by their x and y grid coordinates. Then it makes {sf} geometry objects and calculates the angles between them, and finally does a bit of arithmetic to find if the angles are less than the critical angle. An important aspect to the calculation is that if the angle between two neighbours is greater than 180 degrees, the opposite angle is taken. So if an angle is 275 degrees, the recorded angle is 360 - 275 = 85 degrees. In this example by taking the opposite angle the value of v_{j} changes from 0 (275 gt 90) to 1 (85 lt 90).
The angleCalc() function is:
#' Calculate angle between two sf point objects
#'
#' @param x point feature of class 'sf'
#' @param y point feature of class 'sf'
#'
#' @return azimuthal from x to y, in degrees
#'
#' @examples
#' p1 <- st_point(c(0,1))
#' p2 <- st_point(c(1,2))
#' angleCalc(p1, p2)
#'
#' @export
#'
angleCalc <- function(x, y) {
dst_diff <- as.numeric(x - y)
return((atan2(dst_diff[1], dst_diff[2]) + pi) / 0.01745329)
}
I also wrote some tests for the function to make sure it worked properly. First, packages, all for visualising results, not needed for calculations:
library(ggplot2)
library(patchwork)
library(knitr)
Define a function to create maps to visualise the distribution of neighbours around a focal tree:
linemake <- function(dat) {
out <- data.frame(
xmin = rep(dat$x[1], times = nrow(dat) - 1),
ymin = rep(dat$y[1], times = nrow(dat) - 1),
xmax = dat$x[-1], ymax = dat$y[-1])
return(out)
}
plotmake <- function(dat) {
lout <- linemake(dat)
dat$label <- c("F", "1", "2", "3", "4")
ggplot() +
geom_segment(data = lout,
aes(x = xmin, y = ymin, xend = xmax, yend = ymax)) +
geom_label(data = dat, aes(x = x, y = y, label = label)) +
coord_equal() +
theme_classic() +
labs(x = "X", y = "Y")
}
Create a number of test cases with different spatial distributions of neighbours. Each test case should produce a different value of Wi for the focal tree.
test_list <- list(
test1 = data.frame(x = c(0,0,1,0,-1), y = c(0,1,0,-1,0)),
test2 = data.frame(x = c(0,0,1,0,-0.5), y = c(0,1,0,-1,0.5)),
test3 = data.frame(x = c(0,0,0.5,0,-0.5), y = c(0,1,0.5,-1,0.5)),
test4 = data.frame(x = c(0,0,0.2,-0.4,-0.5), y = c(0,1,0.5,-0.1,0.5)),
test5 = data.frame(x = c(0,0,0.2,-0.5,-0.2), y = c(0,1,0.9,0.4,0.5))
)
Create the maps to visualise layout of the test cases:
plot_list <- lapply(test_list, plotmake)
wrap_plots(plot_list) +
plot_layout(nrow = 1)
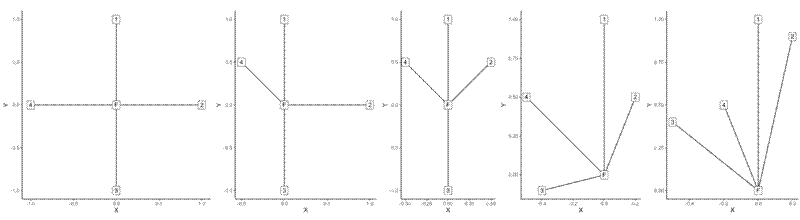
Finally run the winkelmass() function and extract the first value which refers to the focal tree:
kable(
unlist(lapply(test_list, function(x) {
winkelmass(x$x, x$y)[1]
})))
| Ex | Wi |
|---|---|
| test1 | 0.00 |
| test2 | 0.25 |
| test3 | 0.50 |
| test4 | 0.75 |
| test5 | 1.00 |
As you can see, for a given value of k, there are k+1 possible values of Wi depending on the distribution of neighbours.