Voronoi diagrams partition multi-dimensional space into regions, where all positions within a given region are closest to the same known point in the multi-dimensional space. Voronoi diagrams are named after Georgy Voronoy, but can also be called a Dirichlet tessellation (Peter Gustav Lejeune Dirichlet), or Thiessen polygons (Alfred H. Thiessen), as the same thing was implemented in different fields independently.
I discovered voronoi diagrams when researching methods to describe spatial clustering. I wanted a simple single number measure of how clustered together trees in a plot are, so that I could include that metric as a term in a linear model which looked at factors determining woodland canopy complexity.
What is spatial clustering?
It’s useful to think about the distribution of points (spatial point patterns) by looking at three special cases. The first special case is complete spatial randomness (CSR), where all points are located completely independently of one another. The second is total regularity, where all points are an equal distance from all nearest neighbours, as in a regular lattice. The third case is clustering, where points are attracted to each other, rather than repelled. In ecology speak, you could say that spatial regularity is the result of negative density dependence, while spatial clustering is the result of positive density dependence. Regularity and clustering represent opposite ends of a continuum, with CSR in the middle, representing the neutral, zero effect. The continuum is bounded at the end of regularity; given a number of individuals in a space they can only get so far away from their neighbours. At the clustered end of the continuum, I imagine the theoretical limit to clustering would be when all points occupy the same single point in the plot space. The idea of randomness is difficult to comprehend from just one sample. A random distribution could by chance be identical either an even distribution or a clustered distribution. Experiments on randomly generated data will have to be repeated many times to understand the average effect.
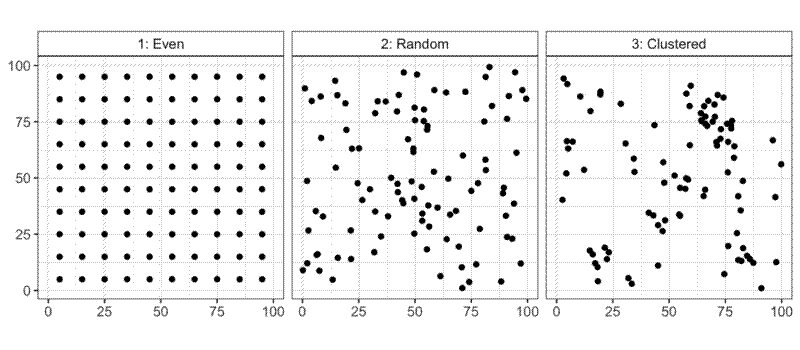
This thought experiment assumes that we are sampling the whole population and the whole available space, but in reality, as the negative density dependence effect increases, the population sampled in the sampling frame merely gets smaller, and the remaining individuals get pushed to outside the sampling frame. Similarly, although there is no theoretical limit on the clustering effect, in biology there is a practical one. It is unlikely that the benefits of clustering among individuals, trees for example, would not be offset by other factors at very close spatial scales, such as the occupation of the same physical space by their trunks.
Extracting metrics from voronoi diagrams
On their own, voronoi diagrams don’t provide any metrics of spatial distribution, they are merely a visual and mathematical representation of a point distribution. From the voronoi polygons however, one can extract various metric related to the size, shape and distribution of the polygons themselves. This is the focus of my investigation in this blog post.
The point distribution norm (Gunzburger and Burkardt 2004 ) is calculated as:
$$ h = \textnormal{max} h_{i} \\ h_{i} = \textnormal{max} |z_{i}-y| $$
where $y$ is the point, and $z_{i}$ are the voronoi cell vertices. $h_{i}$ is the distance from the point to the furthest vertex. Smaller $h$ implies a more uniform distribution.
The point distribution ratio is calculated as:
$$ \mu = \frac{\textnormal{max} h_{i}}{\textnormal{min} h_{i}} $$
where $h_{i}$ is same as above. For a uniform distribution $\mu = 1$.
The regularity measure (Gunzburger and Burkardt 2004) is calculated as:
$$ \chi = \textnormal{max} \chi_{i} \\ \chi_{i} = \frac{2h_{i}}{\gamma_{i}} $$
where $\gamma_{i}$ is the minimum distance between a point $i$ and its nearest neighbour, and where $h_{i}$ is the same as the point distribution norm. For a uniform distribution $\chi = \chi_{i}$, smaller is more uniform.
The cell area deviation (Gunzburger and Burkardt 2004) is calculated as:
$$ \upsilon = \frac{\textnormal{max} V_{i}}{\textnormal{min} V_{i}} $$
where $V_{i}$ is the area of cell $i$. In a uniform distribution, $\upsilon = 1$.
The Polsby-Popper index (Polsby and Popper 1991 ) describes the compactness of a shape. It was originally developed to quantify the degree of gerrymandering of political districts in the USA. For a given voronoi cell, it is calculated as:
$$ PP(D) = \frac{4\pi A(D)}{P(D)^{2}} $$
where $D$ is the voronoi cell, $P(D)$ is the perimeter, and $A(D)$ is the area. Lower values of $PP(D)$ imply less compactness.
Cell elongation is calculated by fitting the minimum bounding rectangle to the voronoi cell, then taking the ratio of the length of the longer side to the shorter side.
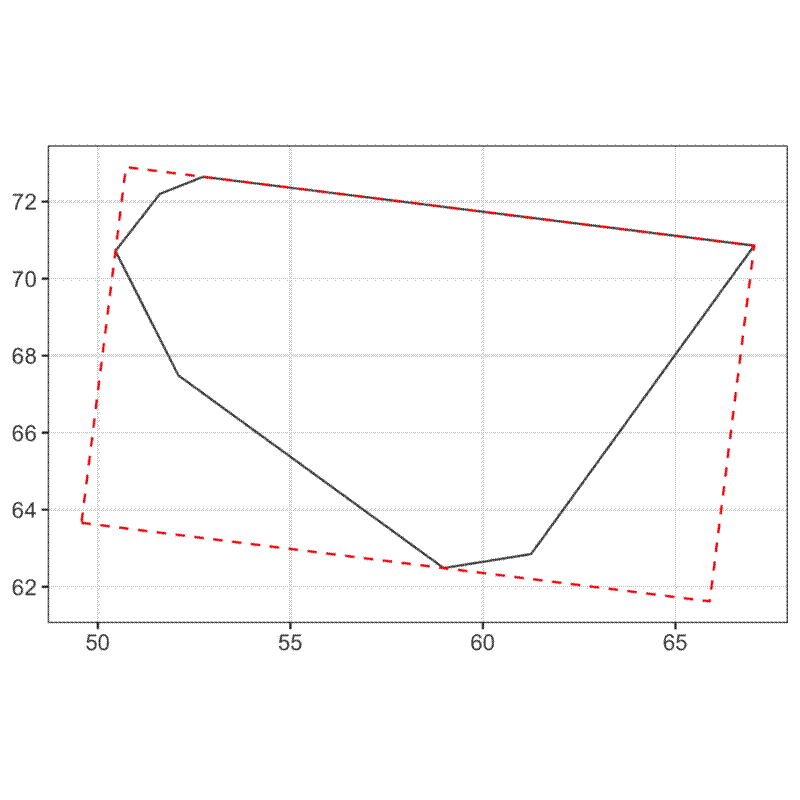
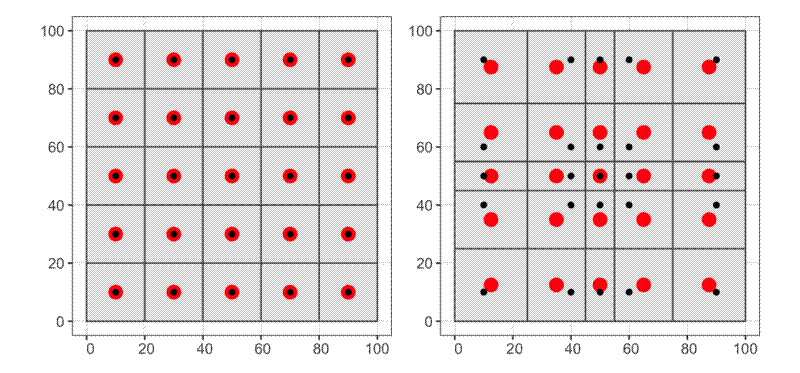
The distance between the centre of gravity (centroid) of the voronoi cell and the point is influenced by the regularity of distribution of neighbouring points. As the distribution deviates from evenness, the distance of points from their cell centroid increases.
Non-voronoi measures of spatial pattern
There are many other measures of spatial point patterns and clustering which aren’t related to voronoi tessellation. I’ll describe some of these methods below, and use them as a comparison to the metrics I derive from the voronoi polygons.
Ripley’s K and Ripley’s L
Ripley’s K function measures the expected number of neighbours found within a given distance of a randomly selected point. To use Ripley’s K to test for CSR, one can simulate a number of random point distributions within the same area as the real population, say 100 or 1000 (multiples of 10 make it easier to estimate P-values), then see whether the real Ripley’s K overlaps with the simulated Ripley’s K functions.
Ripley’s K is often re-calculated as Ripley’s L, also known as the variance stabilised Ripley’s K function. Ripley’s L is approximately constant under CSR. If $L_{obs} < L_{exp}$, the pattern is more regular than expected, while if $L_{obs} > L_{exp}$, the pattern is more clustered than expected. Further Ripley’s L is sometimes plotted as $L_{r} ~ r$, so that the zero line and envelope bounding CSR are horizontal.
Here’s an example of calculating and visualising Ripley’s K, Ripley’s L, and L-r using a real world vegetation survey plot in miombo woodlands in southwest Angola.
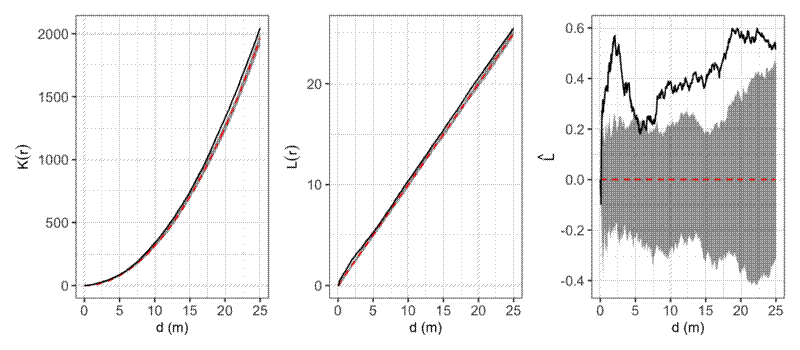
In this example, it looks like stems are generally more clustered than expected at all observed distances. Clustering effects are especially prominent up to ~2.5 m. This may actually be due to some trees having multiple stems growing from the same root stock, but were not observed to connect above the ground, and were thus recorded as separate individuals.
Winkelmass
I wrote about the Uniform Angle Index, or winkelmass, in a previous post. The winkelmass (Pommerening 2006 ) measures the spatial distribution of trees according to the angles between neighbouring trees. For this experiment I calculated the plot level mean and the coefficient of variation of the winkelmass. I won’t go into massive detail about how the winkelmass works, as I’ve talked about it before, but here is a diagram which shows how trees are scored based on the spatial distribution of their four nearest neighbours. Higher scores imply clustering, while lower scores imply regularity.
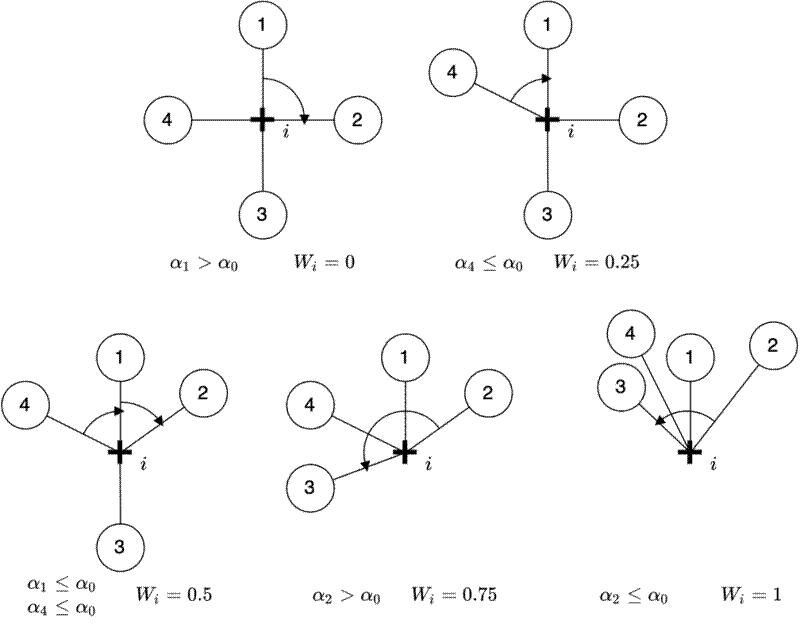
Nearest neighbour distances
Nearest neighbour distance distributions are commonly used to describe spatial point patterns, and are included in the calculations for some of the voronoi-derived metrics. I also chose to calculate the coefficient of variation of nearest neighbour distances as a simple measurement of the heterogeneity in spatial relations among individuals across the plot, and the plot level mean of nearest neighbour distances to measure crowding across the plot.
You can get a visual representation of how close a point distribution is to CSR by comparing the cumulative density distribution of nearest neighbour distances against the theoretical cumulative distribution for a completely random spatial point pattern, which should be $G_{r} = 1 - exp(-\lambda{} * \pi{} * r^{2})$. As with the Ripley’s K and L functions, you can compute simulations to construct a confidence interval ribbon. Points rising above the ribbon are more clustered than expected at that spatial scale.
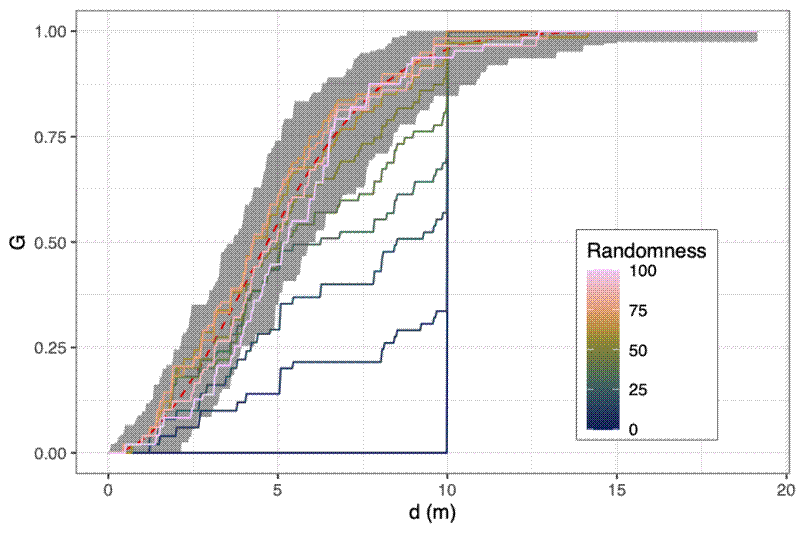
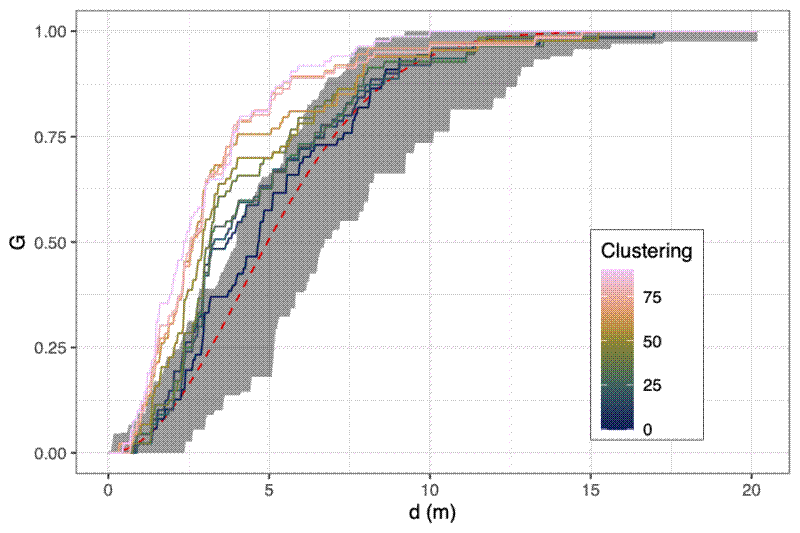
Simulating the behaviour of the metrics
To generate a gradient of evenness to randomness, I started off with a grid of 100 points with 10 metres between each point. Then I sequentially moved randomly chosen points to a random location in the plot. After all points had been moved I assumed the points were distributed completely at random. I repeated this process with 50 replicates. The R code to generate these data is below.
# Create a regular grid of points
# 5 m buffer inside plot
# 10 m between points
# 100x100 m grid
xy_vec <- seq(5, 100, 10)
dat <- expand.grid(xy_vec, xy_vec)
names(dat) <- c("x", "y")
# Number of replicates
reps <- 50
# Replicate grid of points
even_list <- replicate(reps, dat, simplify = FALSE)
# Possible coordinates for movement
c_repls <- seq(0,100,0.1)
# For each replicate, randomly move points
# Move every point once
# Evenness -> randomness
even_csr_points <- mclapply(even_list, function(x) {
# Randomise order in which points are moved
ord <- sample(seq_len(nrow(x)), nrow(x))
# Create list to fill
x <- list(x)
# For each point
for (i in seq_len(nrow(x[[1]]))) {
x[[i + 1]] <- x[[i]]
# Move point to a random location
x[[i + 1]][ord[i], c("x", "y")] <- sample(c_repls, 2, replace = TRUE)
}
return(x)
}, mc.cores = 4)
To generate a gradient of randomness to clustering, I started off with 100 points randomly distributed in the plot. Then I randomly assigned 10 points to act as “cluster points”. For each of the remaining 90 points I then randomly chose points and moved them closer to their nearest cluster point by half their original distance. The R code to generate these data is below.
rand_list <- replicate(reps,
data.frame(
x = sample(c_repls, 100),
y = sample(c_repls, 100)),
simplify = FALSE)
# For each replicate, randomly points closer to nearest cluster point
csr_clust_points <- mclapply(rand_list, function(x) {
# Define points to use as clusters
clust_rows <- sample(seq_len(nrow(x)), 10)
# Extract cluster points
clust_pts <- x[clust_rows,]
# Extract non-cluster points
pts <- x[-clust_rows,]
# Find nearest cluster point for each non-cluster point
nn <- nn2(clust_pts[,1:2], pts[,1:2], k = 1)[[1]]
# Randomise order in which points are moved
ord <- sample(seq_len(nrow(pts)), nrow(pts))
# Create list to fill
pts <- list(pts)
# For each point
for (i in seq_len(nrow(pts[[1]]))) {
pts[[i + 1]] <- pts[[i]]
# Get midpoint between chosen point and cluster point and update coords
pts[[i + 1]][i,1:2] <- (pts[[i + 1]][i,1:2] +
clust_pts[nn[i],1:2]) / 2
}
return(pts)
}, mc.cores = 4)
nn2() comes from the {RANN} package
, which provides a really fast nearest neighbour function. mclapply() comes from the {parallel} package.
For the evenness to randomness dataset, I ended up with 50 replicates, each containing, 101 plots, with each plot containing 100 points. A total of 505000 points for which I had to calculate voronoi polygons. Similarly, the randomness to clustering dataset contained 509190=409500 points. Originally, I was using the st_voronoi() function from the {sf} package to calculate the voronoi polygons, but this was far too slow for my purposes, and the resulting data files were huge, hundreds of MB even when compressed into a .rds file. In the end I settled on using the {deldir} package, which provides a really fast voronoi tessellation function that returns only the coordinates of the vertices of each polygon. This took the size of the data files down to about 2 MB. The R code I used to process the points is below.
vor_polys <- lapply(list(even_csr_points,csr_clust_points), function(i) {
mclapply(seq_along(i), function(x) {
lapply(seq_along(i[[x]]), function(y) {
message(x, ":", y)
vor <- deldir(i[[x]][[y]], rw = c(0,100,0,100))
vor_tiles <- tile.list(vor)
lapply(vor_tiles, function(z) {
cbind(z$x, z$y)
})
})
}, mc.cores = 4)
})
Predictions
In summary, I derived the following metrics of point dispersion:
- Mean winkelmass
- Coefficient of variation of winkelmass
- Coefficient of variation of nearest neighbour distance
- Point distribution norm
- Point distribution ratio
- Regularity measure
- Mean distance between centre of gravity of cell and point
- Coefficient of variation of distance between centre of gravity of cell and point
- Cell area deviation
- Coefficient of variation of cell area
- Coefficient of variation of the Polsby-Popper index
- Coefficient of variation of cell elongation
I thought it would be good practice to make predictions of how each measure might vary along a continuum from evenness to randomness to clustering:
- Mean winkelmass
- Even-random: increase
- Clustered-random: no change
- Coefficient of variation of winkelmass
- Even-random: increase
- Random-clustered: no change
- Coefficient of variation of nearest neighbour distance
- Even-random: increase
- Random-clustered: decrease
- Point distribution norm
- Even-random: increase
- Random-clustered: increase
- Point distribution ratio
- Even-random: increase
- Random-clustered: increase
- Regularity measure
- Even-random: decrease
- Random-clustered: no change
- Mean distance between centre of gravity of cell and point
- Even-random: no change
- Random-clustered: no change
- Coefficient of variation of distance between centre of gravity of cell and point
- Even-random: increase
- Random-clustered: increase
- Cell area deviation
- Even-random: increase
- Random-clustered: increase
- Coefficient of variation of cell area
- Even-random: increase
- Random-clustered: increase
- Coefficient of variation of the Polsby-Popper index
- Even-random: increase
- Random-clustered: increase
- Coefficient of variation of cell elongation
- Even-random: increase
- Random-clustered: increase
Results of simulations
Here are the results for the even-random experiment:
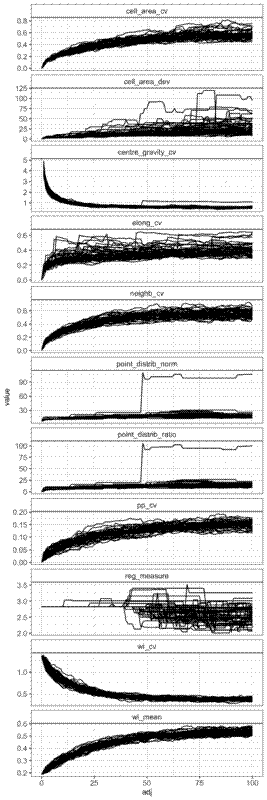
The coefficient of variation of the winkelmass acts in the opposite way to what I expected, decreasing with randomness. I can’t really understanding why this is the case, but it’s a clear trend. I’d expect that as randomness increases, there is greater variation in the spatial distribution of nearest neighbours.
There are also some measures that simply didn’t work. The point distribution norm and point distribution ratio for instance, just didn’t behave how they were suggested in the literature I read. In the plot above you can see that in one simulation the point distribution norm and point distribution ratio suddenly jump up at around 50 substitutions. I looked into what was going on with the that made this happen. It seems that these metrics break down if two points are in exactly the same location. See the plot below, which shows the points and their voronoi cells overlaid at two consecutive intervals. The point which moves between these two intervals is highlighted as a blue square, while the points which don’t move are grey circles. The original position of the points are presented as small markers, and the positions after the movement are presented as large markers. The voronoi cells before the movement are thick black lines, and after the movement are thin orange lines. I guess when a point is moved to exactly the same position as another point their voronoi cells are extremely small and this somehow messes up the metric? I still haven’t got to the bottom of this one.
The regularity measure, although it does show a general decrease in regularity as randomness increases, there is lots of noise, and definitely isn’t monotonic, which I think limits its usefulness. Same with the cell area deviation, it does show a general increase with randomness, but it isn’t consistent.
The coefficient of variation of cell elongation increases with randomness, but it’s quite noisy among simulations, and the measure saturates quickly.
There’s a weird effect where in one simulation the coefficient of variation of the cell centre of gravity suddenly increases before continuing to decrease as in the other simulations. Like with the point distribution norm, this appears to happen when in one simulation a point is moved to the exact same position as another point.
None of the measures increased linearly, but maybe this has more to do with how I generated the transition from evenness to randomness.
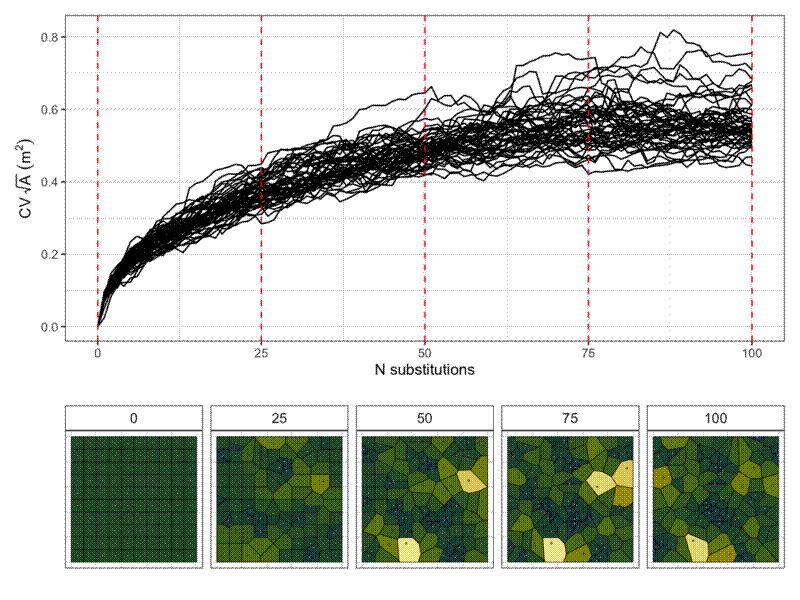
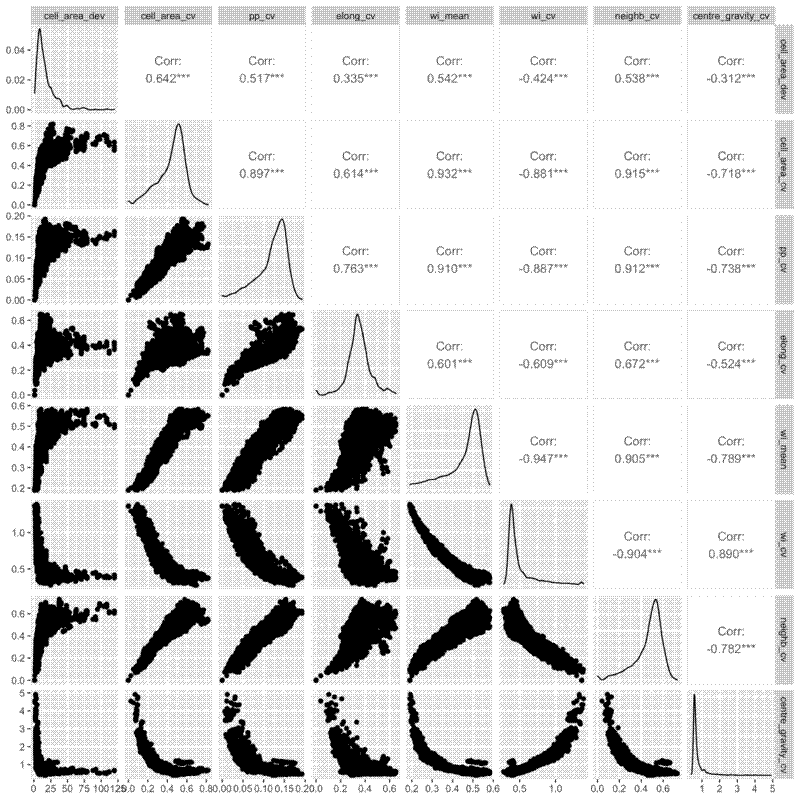
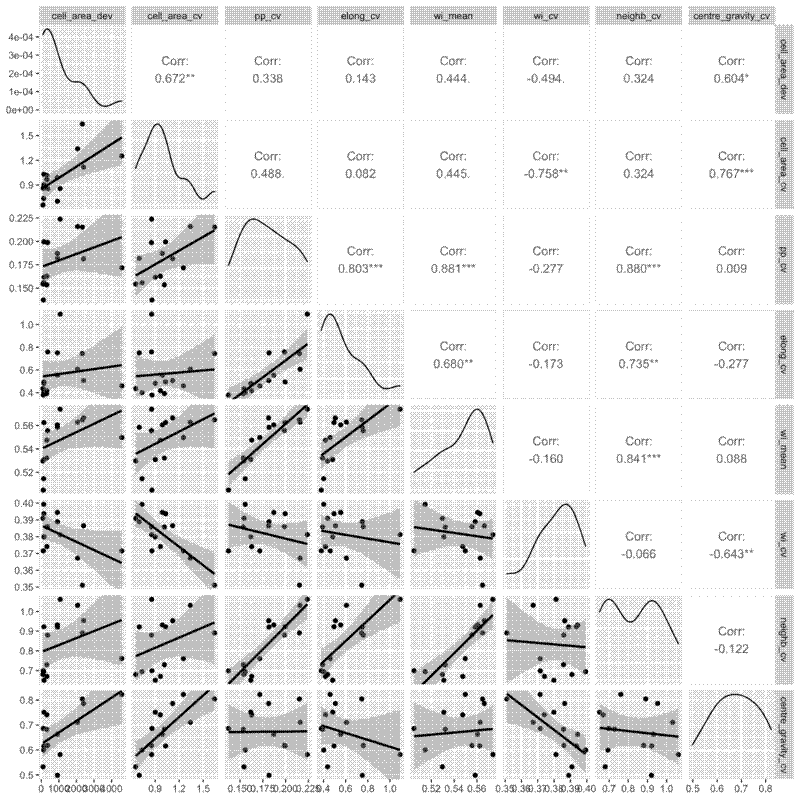
I also did pairwise comparisons of the metrics, to see how they relate to each other. All the metrics were significantly correlated with each other according to a Pearson’s correlation coefficient, but some of the relationships are definitely not linear, so take those correlations with a pinch of salt. I also did a spearmans rank test and there were also all significantly correlated. Some interesting relationsips I see are that as the mean winkelmass increases, the coefficient of variation of the winkelmass decreases. I think this is because as points become less evenly distribution, thus increasing the mean winkelmass, it also means that neighbours are likely to be less evenly distributed, i.e. the points are not totally independent of each other. Also, as cell area coefficient of variation increases, the coefficient of variation of nearest neighbour distances also increases and the coefficient of variation of the polsby-popper index increases. This means basically that cells get longer as they become less evenly distributed, makes sense.
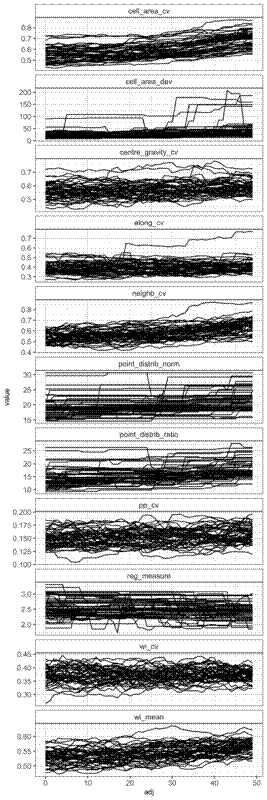
The results for the random-clustered data are not so pretty. Some metrics, like the coefficient of cell area and mean winkelmass do appear to increase with clustering, but understandably, all metrics are heavily dependent on the starting distribution of the points, which were generated randomly. I guess maybe if I had many more simulations I might start to see more clear patterns. It’s curious that some of the measures like the point distribution norm, point distribution ratio, and regularity measure stay the same for many point movements, then suddenly increase or decrease, while other metrics are a lot more sensitive to each movement. This is because some of the metrics, particularly those presented in Gunzburger and Burkardt (2004), rely on changes in the maximum value of some metric measured across all points, so there’s often no observed change in the metrics even though there’s lots of change happening with values less than the maximum.
A real world test
I have some real data from my field site in southwest Angola, Bicuar National Park, where we have 15 permanent vegetation monitoring plots. Each plot is 100x100 m (1 ha). In each plot I have the location of each tree with at least 1 stem >5 cm stem diameter at 1.3 m height (DBH, diameter at breast height). Tree locations were recorded using an XY grid coordinate system, using tape measures.