In a previous post
I showed some of the work I’ve done to extend the functionality of the check_plot_coord() function in the BIOMASS R package
. Since then I have extended the function even more to account for cases where one or more of the plot corners is un-measured, and also cases where the plot is not rectangular (more than four corners). I have also added more checks and messages with feedback for the user. I have renamed the function to plotPolygonFit().
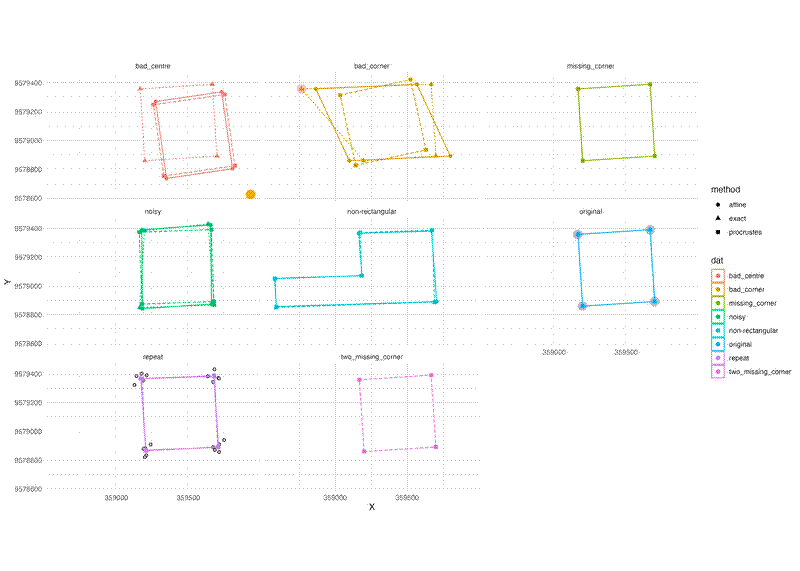
Firstly, here is how the function performs with various different test datasets:
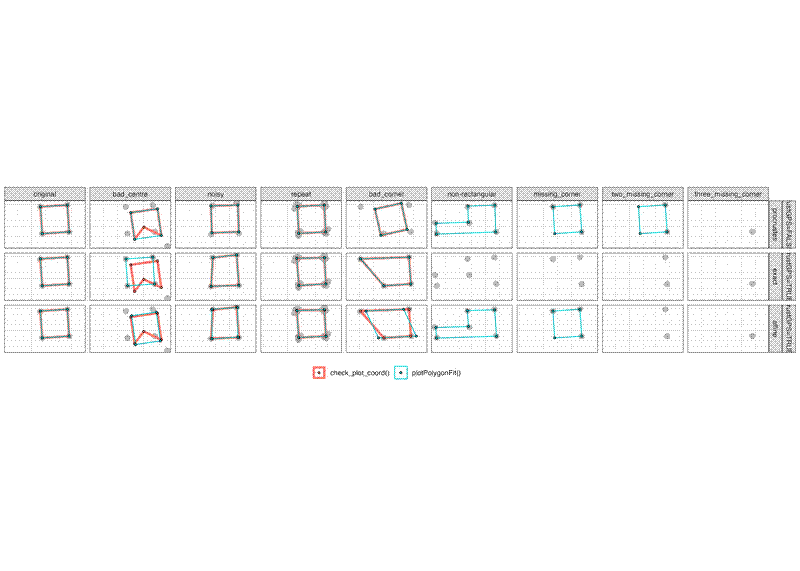
Here is how the function performs in comparison to BIOMASS::check_plot_coord(). Where a panel is missing a polygon it is because either function produces an error:
Below is the code:
#' Fit polygons to plot geo-location points and check locations of trees
#'
#' @description
#' Quality check, geometric fitting, and spatial projection of plot corners and tree coordinates.
#'
#' @details
#' The function maps relative (local/field) coordinates of plot corners to a projected coordinate
#' system. Outlier GPS measurements are identified based on the `max_dist` argument and, if
#' `rm_outliers` is TRUE, the final geometry is recalculated without them.
#'
#' Three transformation methods are available to fit the plot corners and project the trees:
#' * `"exact"`: Corner coordinates are calculated by simply averaging the projected coordinates of
#' multiple GPS measurements for each corner. The projected coordinates of the trees are then
#' calculated from the local coordinates using a bilinear interpolation. This projection strictly
#' requires the local plot to be roughly rectangular (exactly 4 corners).
#' * `"affine"`: An affine transformation is fitted using ordinary least squares (independent scale,
#' rotation, and shear). Trees are projected using the fitted linear models.
#' * `"procrustes"`: A Procrustes analysis is used to preserve the exact shape and dimensions of the
#' plot in the local coordinate system, finding the optimal rotation and translation. Trees are
#' projected applying the resulting rotation matrix and translation vector.
#'
#' If `lonlat_col` is provided instead of `proj_col`, the function will first convert the long/lat
#' coordinates into UTM coordinates. Where plots within a site straddle multiple UTM zones the UTM zone
#' covered by the majority of plot corners will be used, assuming that the extent of the site is small
#' enough to ensure that warping is minimal.
#'
#' If both `lonlat_col` and `proj_col` are provided, only UTM coordinates will be used.
#'
#' When `raster` or `shapefile` are provided, they are cropped/intersected for every plot contained
#' in `point_data` and displayed on in generated plots.
#'
#' @param point_data data frame containing the plot corner coordinates.
#' @param method character string indicating the spatial transformation method to use: `"exact"`, `"affine"`, or `"procrustes"`.
#' @param rel_col character vector of length 2 specifying the column names (resp. x, y) of the corner relative coordinates (that of the field, ie, the local ones).
#' @param proj_col (optional, if `lonlat_col` is not provided) character vector of length 2, specifying the column names (resp. x, y) of the corner projected coordinates.
#' @param lonlat_col (optional, if `proj_col` is not provided) character vector of length 2 specifying the column names of the corner geographic coordinates (long,lat).
#' @param type_col (optional) character indicating the column name in `point_data` that defines the point type (e.g., "corner"). Useful if the dataset contains non-corner reference points.
#' @param plot_col if dealing with multiple plots: a character indicating the variable name for plot IDs in `point_data`.
#' @param tree_data data frame containing the relative coordinates (field/local coordinates) of the trees and optional other tree metrics.
#' @param tree_rel_col character vector specifying the column names of the tree relative coordinates.
#' @param tree_plot_col if dealing with multiple plots: a character indicating the variable name for tree plot IDs in `tree_data`.
#' @param max_dist maximum distance (in meters) above which GPS measurements should be considered outliers (default 10 m).
#' @param rm_outliers if TRUE, outliers are removed from the coordinate calculation of the referenced corners and the geometry is refitted.
#' @param raster filename (character) of the raster to be displayed (typically a CHM raster created from LiDAR data), or a SpatRaster object from terra package.
#' @param shapefile filename (character) of the shapefile to be displayed, or an object of class 'sf' (sf package).
#' @param prop_tree column name variable of `tree_data` for which the tree visualization will be proportional.
#' @param threshold_tree numeric of length 1: the threshold of the `prop_tree` variable at which trees will be displayed on the plot.
#' @param draw_plot logical indicating if the plot design should be displayed and returned.
#' @param ask if TRUE and dealing with multiple plots, prompt user before displaying each plot.
#'
#' @return Returns a list including :
#' - `corner_coord`: a data frame containing the projected coordinates (x_proj and y_proj) and the relative coordinates (x_rel and y_rel) of the corners of the plot
#' - `polygon`: a sf object containing plot's polygon(s)
#' - `plot_design`: if `draw_plot` is TRUE, a ggplot object corresponding to the design of the plot
#' - `outliers`: a data frame containing the projected coordinates and relative coordinates of GPS measurements considered outliers
#' - `UTM_code`: if `lonlat_col` is provided, a character string containing the UTM code of the corner GPS coordinates
#' - `tree_data`: if `tree_data` is provided, a data frame corresponding to tree_data for which the projected coordinates of the trees (x_proj and y_proj) are added, and also a variable telling if the trees are inside the plot (is_in_plot).
#' - `sd_coord`: a data frame containing (for each plot) the average standard deviation of the GPS measurements for each corner.
#'
#' @export
#'
#' @importFrom stats aggregate ave lm predict sd
#' @importFrom sf st_multipoint st_polygon st_sfc st_as_sf st_read st_crs st_transform st_intersection
#' @importFrom ggplot2 ggplot aes geom_point geom_segment geom_polygon geom_path geom_text geom_raster scale_shape_manual scale_color_manual ggtitle theme_minimal theme coord_equal arrow unit element_blank guides guide_legend scale_alpha scale_alpha_manual scale_size scale_fill_gradientn
#' @importFrom terra vect crop as.data.frame rast
#'
#' @author Arthur BAILLY, Arthur PERE, Maxime REJOU-MECHAIN, John L. GODLEE
#'
#' @examples
#' # One plot with repeated measurements of each corner
#' data("NouraguesPlot201")
#' fit_plot201 <- plotPolyFit(
#' point_data = NouraguesPlot201,
#' method = "exact",
#' proj_col = c("Xutm","Yutm"), rel_col = c("Xfield","Yfield"),
#' draw_plot = FALSE)
#' fit_plot201$corner_coord
#' \donttest{
#' fit_plot201$plot_design
#' }
#'
#' # 4 plots with one measurement of each corner
#' data("NouraguesCoords")
#' fit_plots <- plotPolyFit(
#' point_data = NouraguesCoords,
#' method = "affine",
#' proj_col = c("Xutm","Yutm"), rel_col = c("Xfield","Yfield"),
#' plot_col = "Plot", draw_plot = FALSE)
#' fit_plots$corner_coord
#' \donttest{
#' fit_plots$plot_design
#' }
#'
#' # Displaying the associated CHM raster and representing trees proportionally to their diameter
#' plot_204_coords <- NouraguesCoords[NouraguesCoords$Plot==204,]
#' data("NouraguesTrees")
#' plot_204_trees <- NouraguesTrees[NouraguesTrees$Plot == 204, ]
#' nouragues_raster <- terra::rast(
#' system.file("extdata", "NouraguesRaster.tif",
#' package = "BIOMASS", mustWork = TRUE)
#' )
#' fit_plot_204 <- plotPolyFit(
#' point_data = plot_204_coords,
#' method = "procrustes",
#' proj_col = c("Xutm","Yutm"), rel_col = c("Xfield","Yfield"),
#' draw_plot = FALSE,
#' tree_data = plot_204_trees, tree_rel_col = c("Xfield","Yfield"),
#' raster = nouragues_raster, prop_tree = "D", threshold_tree = 25
#' )
#' \donttest{
#' fit_plot_204$plot_design
#' }
#'
plotPolygonFit <- function(point_data,
method = c("exact", "affine", "procrustes"),
rel_col, proj_col = NULL, lonlat_col = NULL, type_col = NULL, plot_col = NULL,
tree_data = NULL, tree_rel_col = NULL, tree_plot_col = NULL,
max_dist = 10, rm_outliers = TRUE,
raster = NULL, shapefile = NULL, prop_tree = NULL, threshold_tree = NULL,
draw_plot = TRUE, ask = TRUE) {
# Rename objects
pdt <- point_data
# Match arguments
method <- match.arg(method)
# Check arguments ------------------------------------------------------------
if (missing(pdt)) {
stop("This function has changed since version 2.2.1. You must provide 'point_data', a dataframe containing point coordinate data")
}
if (!is.data.frame(pdt)) {
stop("'point_data' must be a dataframe or dataframe extension")
}
if (is.null(lonlat_col) & is.null(proj_col)) {
stop("You must provide the column names of at least one set of coordinates: 'lonlat_col' or 'proj_col'")
}
if (!is.null(lonlat_col) & !is.null(proj_col)) {
message("Both 'lonlat_col' and 'proj_col' are specified. 'proj_col' will be used")
lonlat_col <- NULL
}
if (!is.null(proj_col) && !any(proj_col %in% names(pdt))) {
stop("Column names provided by 'proj_col' not found in 'point_data'")
}
if (!is.null(lonlat_col) && !any(lonlat_col %in% names(pdt))) {
stop("Column names provided by 'lonlat_col' not found in 'point_data'")
}
if (!any(rel_col %in% names(pdt))) {
stop("Column names provided by 'rel_col' not found in 'point_data'")
}
if (!is.null(tree_data) && !is.data.frame(tree_data)){
stop("'tree_data' must a dataframe or dataframe extension")
}
if (!is.null(tree_data) && is.null(tree_rel_col)) {
stop("You must provide the column names of the relative coordinates in 'tree_data' using the argument 'tree_rel_col'")
}
if (!is.null(tree_data) && !any(tree_rel_col %in% names(tree_data))) {
stop("Column names provided by 'tree_rel_col' not found in 'tree_data'")
}
if (!is.null(prop_tree) && !any(prop_tree == names(tree_data))) {
stop("Column name provided by 'prop_tree' not found in 'tree_data'")
}
if (!is.null(threshold_tree) && is.null(prop_tree)) {
stop("If 'threshold_tree' is provided, 'prop_tree' must also be provided.")
}
if (!is.null(threshold_tree) && (!is.numeric(threshold_tree) || length(threshold_tree) != 1) ) {
stop("'threshold_tree' must be a numeric of length 1.")
}
if (length(max_dist) != 1) {
stop("'max_dist' must be a numeric of length 1")
}
if (sum(is.na(pdt[, rel_col])) != 0) {
stop("Missing values detected in point relative coordinates.")
}
if (!is.null(proj_col) && method == "exact" && sum(is.na(pdt[, proj_col])) != 0) {
message("Missing values detected in point projected coordinates, these rows will be discarded.")
}
if (!is.null(lonlat_col) && method == "exact" && sum(is.na(pdt[, lonlat_col])) != 0) {
stop("Missing values detected in point longitude/latitude coordinates, these rows will be discarded.")
}
if (!is.null(type_col)) {
cdt_test <- pdt[pdt[[type_col]] == "corner", ]
} else {
cdt_test <- pdt
}
rel_pos <- apply(cdt_test[, rel_col], 1, paste, collapse = ":")
if (method == "exact" && length(unique(rel_pos)) > 4) {
stop("Method 'exact' can only handle exactly four corner points")
}
if (method == "affine" &&
length(unique(rel_pos[!grepl("NA", apply(pdt[, proj_col], 1, paste, collapse = ":"))])) < 3) {
stop("Fewer than three non-missing projected point coordinates, needed for 'affine' method")
}
if (method == "affine" &&
length(unique(rel_pos[!grepl("NA", apply(pdt[, lonlat_col], 1, paste, collapse = ":"))])) < 3) {
stop("Fewer than three non-missing longitude/latitude coordinates, needed for 'affine' method")
}
if (method == "procrustes" &&
length(unique(rel_pos[!grepl("NA", apply(pdt[, proj_col], 1, paste, collapse = ":"))])) < 2) {
stop("Fewer than two non-missing projected point coordinates, needed for 'procrustes' method")
}
if (method == "procrustes" &&
length(unique(rel_pos[!grepl("NA", apply(pdt[, lonlat_col], 1, paste, collapse = ":"))])) < 2) {
stop("Fewer than two non-missing projected point coordinates, needed for 'procrustes' method")
}
if (!is.null(tree_data) && !is.null(plot_col) && is.null(tree_plot_col)) {
stop("Argument 'tree_plot_col' is required if 'plot_col' is provided.")
}
if (!is.null(tree_data) && is.null(plot_col) && !is.null(tree_plot_col)) {
stop("Argument 'plot_col' is required if 'tree_plot_col' is provided.")
}
if (!is.null(plot_col) && !any(plot_col %in% names(pdt))) {
stop("Column name provided by 'plot_col' not found in 'point_data'.")
}
if (!is.null(tree_data) && !is.null(tree_plot_col) && any(!unique(tree_data[[tree_plot_col]]) %in% unique(pdt[[plot_col]])) ) {
message(paste("IDs not matched in 'tree_plot_col' and 'plot_col':" ,
paste(unique(tree_data[[tree_plot_col]])[
!unique(tree_data[[tree_plot_col]]) %in% unique(pdt[[plot_col]])],
collapse = " "), "\n"))
}
if (is.null(type_col)) {
un_pts <- unique(pdt[, c(plot_col, rel_col)])
points_un_plot <- aggregate(un_pts[[rel_col[1]]] ~ un_pts[[plot_col]], FUN = length)
names(points_un_plot) <- c(plot_col, "N")
if (any(points_un_plot$x_rel < 3 | points_un_plot$y_rel < 3)) {
stop("At least three corners per plot required to define a polygon")
}
}
# Data processing ------------------------------------------------------------
# Rename columns
names(pdt)[names(pdt) == rel_col[1]] <- "x_rel"
names(pdt)[names(pdt) == rel_col[2]] <- "y_rel"
if (!is.null(proj_col)) {
names(pdt)[names(pdt) == proj_col[1]] <- "x_proj"
names(pdt)[names(pdt) == proj_col[2]] <- "y_proj"
}
if (!is.null(lonlat_col)) {
names(pdt)[names(pdt) == lonlat_col[1]] <- "lon"
names(pdt)[names(pdt) == lonlat_col[2]] <- "lat"
}
if (!is.null(type_col)) {
names(pdt)[names(pdt) == type_col] <- "type"
}
if (!is.null(plot_col)) {
names(pdt)[names(pdt) == plot_col] <- "plot_ID"
} else {
pdt$plot_ID <- ""
}
# Format tree data
tdt <- NULL
if (!is.null(tree_data)) {
tdt <- tree_data
names(tdt)[names(tdt) == tree_rel_col[1]] <- "x_rel"
names(tdt)[names(tdt) == tree_rel_col[2]] <- "y_rel"
if (!is.null(tree_plot_col)) {
names(tdt)[names(tdt) == tree_plot_col] <- "plot_ID"
} else {
tdt$plot_ID <- ""
}
}
# Transform lat-lon coordinates to UTM if provided
if (!is.null(lonlat_col)) {
UTM_code <- getUTM(pdt$lon, pdt$lat, type = "proj")
if (length(unique(UTM_code)) > 1) {
message("More than one UTM zone detected. Latitude-longitude coordinates will be transformed to the most frequent UTM zone across plots.")
}
utm_unique <- names(sort(table(UTM_code), decreasing = TRUE))[1]
proj_xy <- proj4::project(pdt[, c("lon", "lat")], proj = utm_unique)
pdt$x_proj <- proj_xy$x
pdt$y_proj <- proj_xy$y
}
# Calculate sd_coord
sd_x <- aggregate(x_proj ~ plot_ID + x_rel + y_rel, data = pdt, FUN = sd, na.action = na.pass)
sd_y <- aggregate(y_proj ~ plot_ID + x_rel + y_rel, data = pdt, FUN = sd, na.action = na.pass)
sd_merged <- merge(sd_x, sd_y, by = c("plot_ID", "x_rel", "y_rel"), all = TRUE)
sd_merged$sd_coord <- rowMeans(sd_merged[, c("x_proj", "y_proj")], na.rm = TRUE)
sd_coord <- aggregate(
x = list(sd_coord = sd_merged$sd_coord),
by = list(plot_ID = sd_merged$plot_ID),
FUN = function(x) mean(x, na.rm = TRUE)
)
# Check corner coordinates and calculate tree projected coordinates --------
# Fit models, calculate predictions, and identify outliers (Vectorized & split approach)
apply_fit <- function(df, meth) {
res_list <- lapply(split(df, df$plot_ID), function(sub_dt) {
if (meth == "exact") {
sub_dt$x_proj_pred <- ave(sub_dt$x_proj, sub_dt$x_rel, sub_dt$y_rel, FUN = function(x) mean(x, na.rm = TRUE))
sub_dt$y_proj_pred <- ave(sub_dt$y_proj, sub_dt$x_rel, sub_dt$y_rel, FUN = function(x) mean(x, na.rm = TRUE))
} else if (meth == "affine") {
m_x <- lm(x_proj ~ x_rel + y_rel, data = sub_dt)
m_y <- lm(y_proj ~ x_rel + y_rel, data = sub_dt)
sub_dt$x_proj_pred <- predict(m_x, sub_dt)
sub_dt$y_proj_pred <- predict(m_y, sub_dt)
} else if (meth == "procrustes") {
proc <- BIOMASS:::procrust(
sub_dt[!is.na(sub_dt$x_proj) & !is.na(sub_dt$y_proj), c("x_proj", "y_proj")],
sub_dt[!is.na(sub_dt$x_proj) & !is.na(sub_dt$y_proj), c("x_rel", "y_rel")])
rot <- as.matrix(sub_dt[, c("x_rel", "y_rel")]) %*% proc$rotation
sub_dt$x_proj_pred <- rot[, 1] + proc$translation[1]
sub_dt$y_proj_pred <- rot[, 2] + proc$translation[2]
}
return(sub_dt)
})
do.call(rbind, unname(res_list))
}
pdt <- apply_fit(pdt, method)
# Identify and extract outliers
pdt$outlier <- sqrt((pdt$x_proj - pdt$x_proj_pred)^2 + (pdt$y_proj - pdt$y_proj_pred)^2) > max_dist
pdt$n_meas <- ave(rep(1, nrow(pdt)), pdt$plot_ID, pdt$x_rel, pdt$y_rel, FUN = length)
pdt$n_outliers <- ave(as.integer(pdt$outlier), pdt$plot_ID, pdt$x_rel, pdt$y_rel, FUN = sum)
outliers_df <- pdt[pdt$outlier == TRUE & !is.na(pdt$outlier), c("plot_ID", "x_proj", "y_proj", "x_rel", "y_rel")]
if (rm_outliers) {
pdt <- pdt[!(pdt$outlier == TRUE & !is.na(pdt$outlier) & pdt$n_meas > pdt$n_outliers), ]
pdt <- apply_fit(pdt, method)
}
# Extract corners and aggregate to 1 row per unique corner
if (!is.null(type_col)) {
cdt <- pdt[pdt$type == "corner", ]
} else {
cdt <- pdt
}
if (method == "exact" & any(is.na(cdt[,c("x_proj_pred", "y_proj_pred")]))) {
stop("No valid location for one or more corners")
}
corner_checked <- aggregate(cbind(x_proj_pred, y_proj_pred) ~ plot_ID + x_rel + y_rel,
data = cdt, FUN = mean, na.rm = TRUE)
names(corner_checked)[
names(corner_checked) %in% c("x_proj_pred", "y_proj_pred")] <- c("x_proj", "y_proj")
# Sort corners counter-clockwise radially or by order in dataframe
if (nrow(cdt) <= 4) {
corner_checked <- do.call(rbind, lapply(split(corner_checked, corner_checked$plot_ID), function(sub_c) {
centroid_x <- mean(sub_c$x_rel)
centroid_y <- mean(sub_c$y_rel)
angles <- atan2(sub_c$y_rel - centroid_y, sub_c$x_rel - centroid_x)
sub_c[order(angles), ]
}))
} else {
cdt$pos <- seq_len(nrow(cdt))
orig_ord <- aggregate(pos ~ plot_ID + x_rel + y_rel, data = cdt, FUN = min)
corner_checked <- merge(corner_checked, orig_ord, by = c("plot_ID", "x_rel", "y_rel"))
corner_checked <- corner_checked[order(corner_checked$plot_ID, corner_checked$pos), ]
corner_checked$pos <- NULL # Clean up
}
rownames(corner_checked) <- NULL
# Tree projection ------------------------------------------------------------
if (!is.null(tree_data)) {
tdt_list <- lapply(split(tdt, tdt$plot_ID), function(sub_tdt) {
pid <- sub_tdt$plot_ID[1]
sub_corners <- corner_checked[corner_checked$plot_ID == pid, ]
if (method == "exact") {
x_A <- sub_corners$x_rel[1]; y_A <- sub_corners$y_rel[1]
x_B <- sub_corners$x_rel[2]; y_B <- sub_corners$y_rel[2]
x_C <- sub_corners$x_rel[3]; y_C <- sub_corners$y_rel[3]
x_D <- sub_corners$x_rel[4]; y_D <- sub_corners$y_rel[4]
u_A <- sub_corners$x_proj[1]; v_A <- sub_corners$y_proj[1]
u_B <- sub_corners$x_proj[2]; v_B <- sub_corners$y_proj[2]
u_C <- sub_corners$x_proj[3]; v_C <- sub_corners$y_proj[3]
u_D <- sub_corners$x_proj[4]; v_D <- sub_corners$y_proj[4]
x <- sub_tdt$x_rel
y <- sub_tdt$y_rel
rate_A <- (1 - (x - x_A) / (x_C - x_A)) * (1 - (y - y_A) / (y_C - y_A))
rate_B <- (1 - (x - x_B) / (x_D - x_B)) * (1 - (y - y_B) / (y_D - y_B))
rate_C <- (1 - (x - x_C) / (x_A - x_C)) * (1 - (y - y_C) / (y_A - y_C))
rate_D <- (1 - (x - x_D) / (x_B - x_D)) * (1 - (y - y_D) / (y_B - y_D))
sub_tdt$x_proj <- rate_A * u_A + rate_B * u_B + rate_C * u_C + rate_D * u_D
sub_tdt$y_proj <- rate_A * v_A + rate_B * v_B + rate_C * v_C + rate_D * v_D
} else if (method == "affine") {
m_x <- lm(x_proj ~ x_rel + y_rel, data = sub_corners)
m_y <- lm(y_proj ~ x_rel + y_rel, data = sub_corners)
sub_tdt$x_proj <- predict(m_x, sub_tdt)
sub_tdt$y_proj <- predict(m_y, sub_tdt)
} else {
proc <- BIOMASS:::procrust(sub_corners[, c("x_proj", "y_proj")], sub_corners[, c("x_rel", "y_rel")])
rot <- as.matrix(sub_tdt[, c("x_rel", "y_rel")]) %*% proc$rotation
sub_tdt$x_proj <- rot[, 1] + proc$translation[1]
sub_tdt$y_proj <- rot[, 2] + proc$translation[2]
}
# Build a temporary local polygon and check if tree points fall within it
poly_mat_rel <- as.matrix(rbind(sub_corners[, c("x_rel", "y_rel")], sub_corners[1, c("x_rel", "y_rel")]))
poly_sf_rel <- sf::st_sfc(sf::st_polygon(list(poly_mat_rel)))
tree_sf_rel <- sf::st_as_sf(sub_tdt, coords = c("x_rel", "y_rel"))
sub_tdt$is_in_plot <- as.logical(sf::st_intersects(tree_sf_rel, poly_sf_rel, sparse = FALSE))
return(sub_tdt)
})
tdt <- do.call(rbind, unname(tdt_list))
}
# Create SF polygons ------------------------------------------------------
poly_list <- list()
for (pid in unique(corner_checked$plot_ID)) {
sub_corners <- corner_checked[corner_checked$plot_ID == pid, ]
poly_mat <- as.matrix(rbind(sub_corners[, c("x_proj", "y_proj")], sub_corners[1, c("x_proj", "y_proj")]))
poly_geom <- st_polygon(list(poly_mat))
# Conditionally create the geometry column to avoid NA type errors in sf
if (!is.null(lonlat_col)) {
geom_sfc <- st_sfc(poly_geom, crs = utm_unique)
} else {
geom_sfc <- st_sfc(poly_geom)
}
poly_list[[pid]] <- st_sf(plot_ID = pid, geometry = geom_sfc)
}
if (length(poly_list) > 0) {
corner_polygon <- do.call(rbind, poly_list)
} else {
corner_polygon <- NULL
}
# Plotting ------------------------------------------------------
plot_list <- list()
if (draw_plot) {
if (!is.null(raster) && is.character(raster)) raster <- terra::rast(raster)
if (!is.null(shapefile) && is.character(shapefile)) shapefile <- st_read(shapefile, quiet=TRUE)
for (pid in unique(corner_checked$plot_ID)) {
sub_corners <- corner_checked[corner_checked$plot_ID == pid, ]
poly_sf <- corner_polygon[corner_polygon$plot_ID == pid, ]
p <- ggplot()
if (!is.null(raster) && inherits(raster, "SpatRaster")) {
r_crop <- as.data.frame(crop(raster, vect(poly_sf), mask = TRUE), xy = TRUE)
if (nrow(r_crop) > 0) {
p <- p +
geom_raster(data = r_crop, aes(x = x, y = y, fill = .data[[names(r_crop)[3]]])) +
scale_fill_gradientn(colours = rev(terrain.colors(10)))
}
}
if (!is.null(shapefile)) {
# Perform the intersection using the matching CRS
if (st_crs(shapefile) != st_crs(poly_sf)) {
shp_tmp <- st_transform(shapefile, st_crs(poly_sf))
} else {
shp_tmp <- shapefile
}
shp_crop <- st_intersection(shp_tmp, poly_sf)
if (nrow(shp_crop) > 0) {
p <- p + geom_sf(data = shp_crop, fill = NA, color = "darkgreen")
}
}
# Extract the original (pre-fitted) corner measurements for this plot
sub_orig <- cdt[cdt$plot_ID == pid, ]
poly_df <- rbind(sub_corners, sub_corners[1, ])
p <- p +
geom_path(data = poly_df, aes(x = x_proj, y = y_proj), linewidth = 1.2, color = "black") +
# Layer 1: Original GPS measurements (Open Triangles)
geom_point(data = sub_orig, aes(x = x_proj, y = y_proj), shape = 2, size = 2, color = "black") +
# Layer 2: Final Fitted Corners (Filled Squares)
geom_point(data = sub_corners, aes(x = x_proj, y = y_proj), shape = 15, size = 3, color = "black")
# Add blue orientation arrows for the local grid (x_rel, y_rel)
if (nrow(sub_corners) >= 3) {
arrow_df <- data.frame(
x = sub_corners$x_proj[1],
y = sub_corners$y_proj[1],
xend = c(
sub_corners$x_proj[1] + (sub_corners$x_proj[2] - sub_corners$x_proj[1]) / 4,
sub_corners$x_proj[1] + (sub_corners$x_proj[nrow(sub_corners)] - sub_corners$x_proj[1]) / 4
),
yend = c(
sub_corners$y_proj[1] + (sub_corners$y_proj[2] - sub_corners$y_proj[1]) / 4,
sub_corners$y_proj[1] + (sub_corners$y_proj[nrow(sub_corners)] - sub_corners$y_proj[1]) / 4
),
label = c("x_rel", "y_rel")
)
p <- p +
geom_segment(data = arrow_df, aes(x = x, y = y, xend = xend, yend = yend),
arrow = arrow(length = unit(0.3, "cm")), color = "blue", linewidth = 1) +
geom_text(data = arrow_df, aes(x = xend, y = yend, label = label),
color = "blue", vjust = -1, hjust = -0.1)
}
sub_outliers <- outliers_df[outliers_df$plot_ID == pid, ]
if (nrow(sub_outliers) > 0) {
p <- p + geom_point(data = sub_outliers, aes(x = x_proj, y = y_proj), shape = 4, size = 3, color = "red")
}
if (!is.null(tdt)) {
sub_trees <- tdt[tdt$plot_ID == pid, ]
# Only attempt to plot trees and build scales if this plot actually contains trees
if (nrow(sub_trees) > 0) {
# Clean character column for the legend
sub_trees$Location <- ifelse(sub_trees$is_in_plot, "Inside Plot", "Outside Plot")
if (!is.null(prop_tree)) {
if (!is.null(threshold_tree)) {
filt <- sub_trees[[prop_tree]] >= threshold_tree
} else {
filt <- rep(TRUE, nrow(sub_trees))
}
p <- p +
geom_point(data = sub_trees[sub_trees$is_in_plot == TRUE & filt, ],
aes(x = x_proj, y = y_proj, size = .data[[prop_tree]], alpha = .data[[prop_tree]]), shape = 1) +
geom_point(data = sub_trees[sub_trees$is_in_plot == FALSE, ], aes(x = x_proj, y = y_proj), shape = 13, size = 2) +
scale_alpha(range = c(0.2, 1)) +
scale_size(range = c(0.1, 6))
} else {
p <- p +
geom_point(data = sub_trees, aes(x = x_proj, y = y_proj, shape = Location), alpha = 0.6) +
scale_shape_manual(values = c("Inside Plot" = 1, "Outside Plot" = 13), name = "Tree Location")
}
}
}
p <- p +
coord_equal() +
theme_minimal() +
ggtitle(paste("Plot:", pid))
plot_list[[pid]] <- p
if (ask) {
print(p)
if (pid != unique(corner_checked$plot_ID)[length(unique(corner_checked$plot_ID))]) {
readline(prompt = "Press [enter] to see the next plot")
}
} else {
print(p)
}
}
}
# Back-transform UTM to longlat -------------------------------
if (!is.null(lonlat_col) && exists("utm_unique")) {
ll_corners <- proj4::project(xy = corner_checked[, c("x_proj", "y_proj")], proj = utm_unique, inverse = TRUE)
corner_checked$long <- ll_corners$x
corner_checked$lat <- ll_corners$y
if (!is.null(tdt)) {
ll_trees <- proj4::project(xy = tdt[, c("x_proj", "y_proj")], proj = utm_unique, inverse = TRUE)
tdt$long <- ll_trees$x
tdt$lat <- ll_trees$y
}
}
# Clean returns -----------------------------------------------------------
if (all(unique(corner_checked$plot_ID) == "")) {
corner_checked$plot_ID <- NULL
}
output <- list(
corner_coord = corner_checked,
polygon = corner_polygon
)
if (draw_plot) {
if (length(plot_list) == 1) {
output$plot_design <- plot_list[[1]]
} else {
output$plot_design <- plot_list
}
}
if (nrow(outliers_df) > 0) {
output$outliers <- outliers_df
}
if (!is.null(lonlat_col)) {
output$UTM_code <- utm_unique
}
if (!is.null(tree_data)) {
if (all(unique(tdt$plot_ID) == "")) {
tdt$plot_ID <- NULL
}
output$tree_data <- tdt
}
# Clean up sd_coord formatting based on plot counts
if (nrow(sd_coord) == 1 ) {
output$sd_coord <- sd_coord$sd_coord
} else {
output$sd_coord <- sd_coord
}
# Return
return(output)
}